English version /
English version /
 Version française
Version française
 Download .zip
Download .zip
 View on GitHub
View on GitHub
 View on GitLab IN2P3
View on GitLab IN2P3
PyCoA (Python Covid Analysis) is a Python™ code framework which provides:
- a simple access to the main databases of Covid19;
- tools to display and analyse data related to Covid19, such as time series or maps.
This environment is designed to be accessible to non-specialists: high-school students who are learning Python™, university students, data journalists, even scientists who are not particularly familiar with extracting data from online databases. Simple analyses can be readily performed, and deeper ones may be programmed by people with a reasonable Python™ proficiency.
The PyCoA tool provides access to several databases and reformats its data. PyCoA transparently joins the data with geo-localization databases (management of country or region names, creation of maps). This geolocation information can also be used for other applications outside the Covid19 problematics.
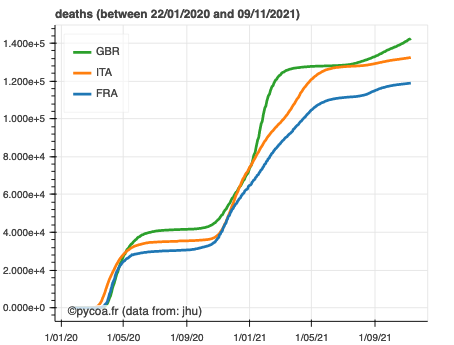
The PyCoA tool is designed to be used in a jupyter environment, installed locally or on a remote server (like Google colaboratory or Binder). This simplifies installation and ensures, thanks to the Bokeh library, powerful graphical outputs with very few lines of code for the user, as the following examples attest.
import coa.front as cf
cf.plot(where=['France', 'Italy', 'United kingdom'])
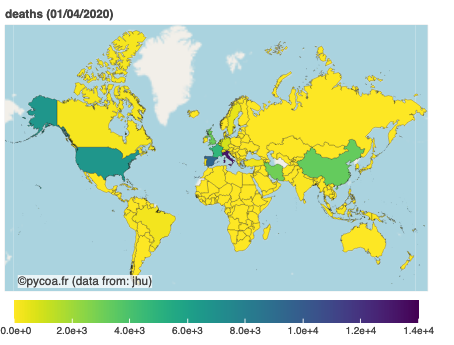
cf.map(where='world',what='daily',when='01/04/2020')
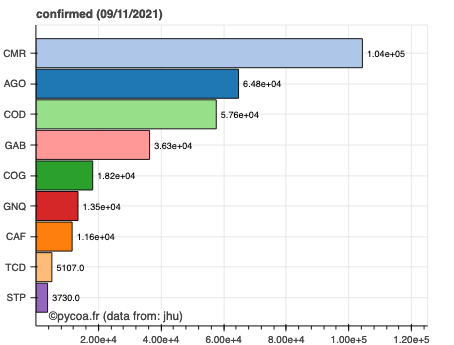
cf.hist(where='middle africa', which='tot_confirmed',what='cumul')
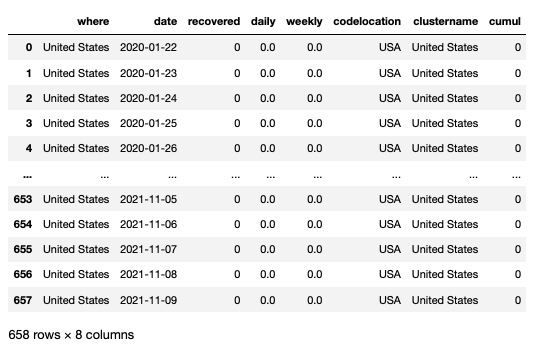
cf.get(where='usa', what='daily', which='tot_recovered',output='pandas')
About us
Physicists on particle physics experiments at CERN or studying complex matter, used to managing big data, we wished to share our skills in statistics and data management to the largest number of people interested in the analysis of data related to the Covid19 pandemic. We believe that data mining and statistics should help everyone to better understand one of the most important phenomena in recent history.
This is the reason why we have initiated the PyCoA project, which stands for Python™ Covid19 Analysis. This tool that can be used online, with a simple interface and clear modelling schemes. It is available online through Google Colaboratory or Binder, a Python™ notebook infrastructure, with no installation effort. Collaborative work can be therefore performed, with access to a huge computational infrastructure (like GPUs for deep learning analyses of Covid19 data).
Origins
The PyCoA project, initially named CoCoA, was born from the participation in April 2020 to the hackathon organized by UltraHack.org.
Selected in final phase, the project is free and open source. It has continued to evolve since then and underwent a major update during the Hackathon Covid19 organized by the Direction interministérielle de la transformation publique in April 2021.
Its code is public as well as the notebooks and are used as references or examples of use during animations or workshops (during the Salon Culture et Jeux Mathématiques 2021 or during the Fête de la Science 2021).
PyCoA: generic software for numerical analysis in Python.
The project was originally developed for epidemiological studies related to Covid19. However, it can be used for other types of studies.
Any analyses involving time series data associated with numerical and geographical variables can use PyCoA. Studies will be greatly simplified with clear and precise graphical representations.
License
The PyCoA project is under MIT license.
Authors
- Tristan Beau - Université de Paris - LPNHE laboratory
- Julien Browaeys - Université de Paris - MSC laboratory
- Olivier Dadoun - CNRS/in2p3 - LPNHE laboratory
Institutions





We talk about us…
- On the MODCOV19 (Modelisation and Covid-19) website from CNRS : PyCoa : Python Covid analysis
- In the Acteurs Publics L’héritage du “Hackathon Covid” passé à la loupe
- In the news of LPNHE - Laboratoire de Physique Nucléaire et Hautes Énergies (july 2021) : Pycoa, un logiciel pour mieux comprendre la pandémie due à la Covid-19
- In the news of Sorbonne Université (july 2021) : PyCoa : un logiciel gratuit d’analyse des données de la Covid-19
- In the news of the Physics departement of Université de Paris (july 2021) : «Un logiciel pour mieux comprendre la pandémie»
- In the news of Université de Paris (june 2021) : «Un logiciel pour mieux comprendre la pandémie»
- At the 22ème salon culture et jeux mathématiques (may 2021) :
https://salon-math.fr/2021/04/14/pycoa/ - At the hackathon covid - lutter ensemble (april 2021) :
https://forum.hackathon-covid.fr/t/3a-pycoa-une-analyse-python-des-donnees-covid-pour-tous/251